Genetic Code
- It is the sequence of nucleotides (nitrogen bases) in mRNA that contains information for protein synthesis (translation).
- The sequence of 3 bases determining a single amino acid is called codon.
- George Gamow suggested that for coding 20 amino acids, the code should be made up of 3 nucleotides. Thus, there are 64 codons (43 = 4 × 4 × 4).
- Har Gobind Khorana developed the chemical method in synthesizing RNA molecules with defined combinations of bases (homopolymers & copolymers).
- Marshall Nirenberg developed a cell-free system for protein synthesis.
- Severo Ochoa (polynucleotide phosphorylase enzyme) is used to polymerize RNA with defined sequences in a template-independent manner.
20 types of amino acids involved in translation
- Alanine (Ala)
- Arginine (Arg)
- Asparagine (Asn)
- Aspartic acid (Asp)
- Cysteine (Cys)
- Glutamine (Gln)
- Glutamic acid (Glu)
- Glycine (Gly)
- Histidine (His)
- Isoleucine (Ile)
- Leucine (Leu)
- Lysine (Lys)
- Methionine (Met)
- Phenylalanine (Phe)
- Proline (Pro)
- Serine (Ser)
- Threonine (Thr)
- Tryptophan (Trp)
- Tyrosine (Tyr)
- Valine (Val)
The codons for various amino acids
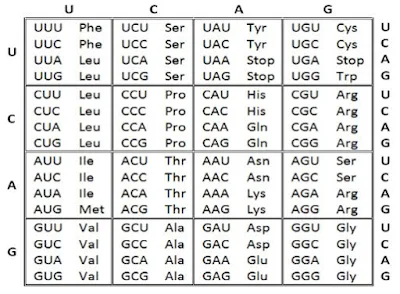
The codons for various amino acids
Salient features of genetic code
- Codon is triplet (three-letter code).
- 61 codons code for amino acids. 3 codons (UAA, UAG & UGA) do not code for any amino acids. They act as stop codons (termination codons or non-sense codons).
- Genetic code is universal. E.g., from bacteria to human, UUU codes for Phenylalanine. Some exceptions are found in mitochondrial codons and in some protozoans.
- No punctuations between adjacent codons (comma-less code). The codon is read in mRNA in a contiguous fashion.
- Genetic code is non-overlapping.
- An amino acid is coded by more than one codon (except AUG for methionine & UGG for tryptophan). Such codons are called degenerate codons.
- Genetic code is unambiguous and specific, i.e., one codon specifies only one amino acid.
- AUG has dual functions. It codes for Methionine and acts as an initiator codon. In eukaryotes, methionine is the first amino acid, and formyl methionine in prokaryotes.
Mutations and Genetic Code
- Relationship between genes & DNA is best understood by mutation studies. Deletions & rearrangements in DNA may cause loss or gain of a gene and thus a function.
- Insertion or deletion of one or two bases changes the reading frame from the point of insertion or deletion. It is called frame-shift insertion or deletion mutations.
- Insertion/deletion of three or its multiple bases inserts or deletes one or multiple codons. The reading frame remains unaltered from that point onwards. Hence, one or multiple amino acids are inserted/deleted.
- It proves that a codon is a triplet and is read contiguously.
Types of RNA
- mRNA (messenger RNA): Provides the template for translation (protein synthesis).
- rRNA (ribosomal RNA): Plays a structural & catalytic role during translation. E.g., 23S rRNA in bacteria acts as a ribozyme.
- tRNA (transfer RNA or sRNA or soluble RNA): Brings amino acids for protein synthesis and reads the genetic code.
- Francis Crick postulated the presence of an adapter molecule that can read the code and link with amino acids.
- tRNA is called an adapter molecule because it has:
- An Anticodon (NODOC) loop that has bases complementary to the codon.
- An amino acid acceptor end to which the amino acid binds.
- Ribosome binding loop.
- Enzyme binding loop.

Structure of tRNA
- For initiation, there is another tRNA called initiator tRNA.
- There are no tRNAs for stop codons.
- Secondary (2-D) structure of tRNA looks like a clover-leaf. The 3-D structure looks like an inverted ‘L’.
Select Your Topic 👇
-
Topic 1: The DNA
Topic 2: The Search for Genetic Material
Topic 3: Properties of Genetic Material, RNA World
Topic 4: DNA Replication
Topic 5: Transcription
Topic 6: Genetic Code, Types of RNA
Topic 7: Translation (Protein Synthesis)
Topic 8: Regulation of Gene Expression, Operon Concept
Topic 9: Human Genome Project (HGP)
Topic 10: DNA Fingerprinting
